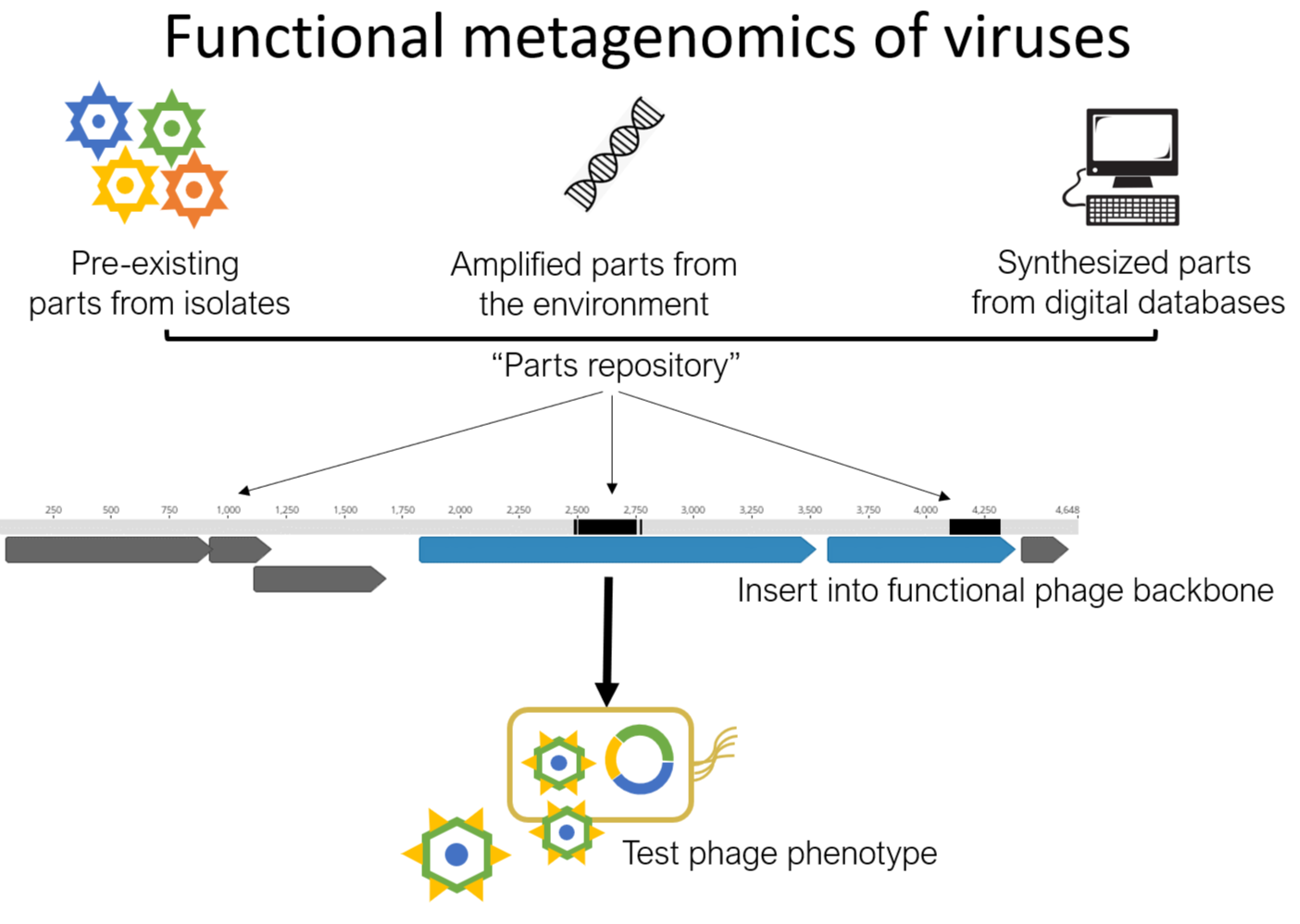
Our main experimental system is the enterobacterial phage genus Enterogokushovirus, a member of the Gokushovirinae subfamily of the Microviridae. These small, single-stranded DNA phages are found everywhere from the human gut to deep sea trenches, but with few exceptions are poorly understood.
What makes this system so unique is that it is amendable to genetic manipulation using simple molecular biological techniques – genes or entire genomes can be synthesized in-vitro and functional viruses can be produced essentially from scratch. Hybrid phages can be built with parts sourced from isolates, from environmental PCR or de-novo synthesis from digital templates. Therefore, the function of individual genomic parts can be studied in great detail.
Our long-term research goal is to gain a functional understanding of microvirus genetics that allows us to predict a given virus’ interaction with its host based on genomic information, and to build microvirus genomes and functional viruses to target specific bacterial strains. Ultimately, we want to develop microviruses as a modular tool for phage therapy and engineering microbiomes by selected killing or culling of undesirable microorganisms.
However, our system is brand new – most aspects of its biology are yet to be determined, providing ample opportunities for a wide variety of research projects. In the short term, we want to understand how different regions in the phage genome can be exchanged or modified to alter the phage’s ability to infect bacteria or protect bacterial hosts from other phages. We also want to expand our repertoire of microviruses using classic and novel isolation techniques, and are further open to the study of any other atpyical phage.
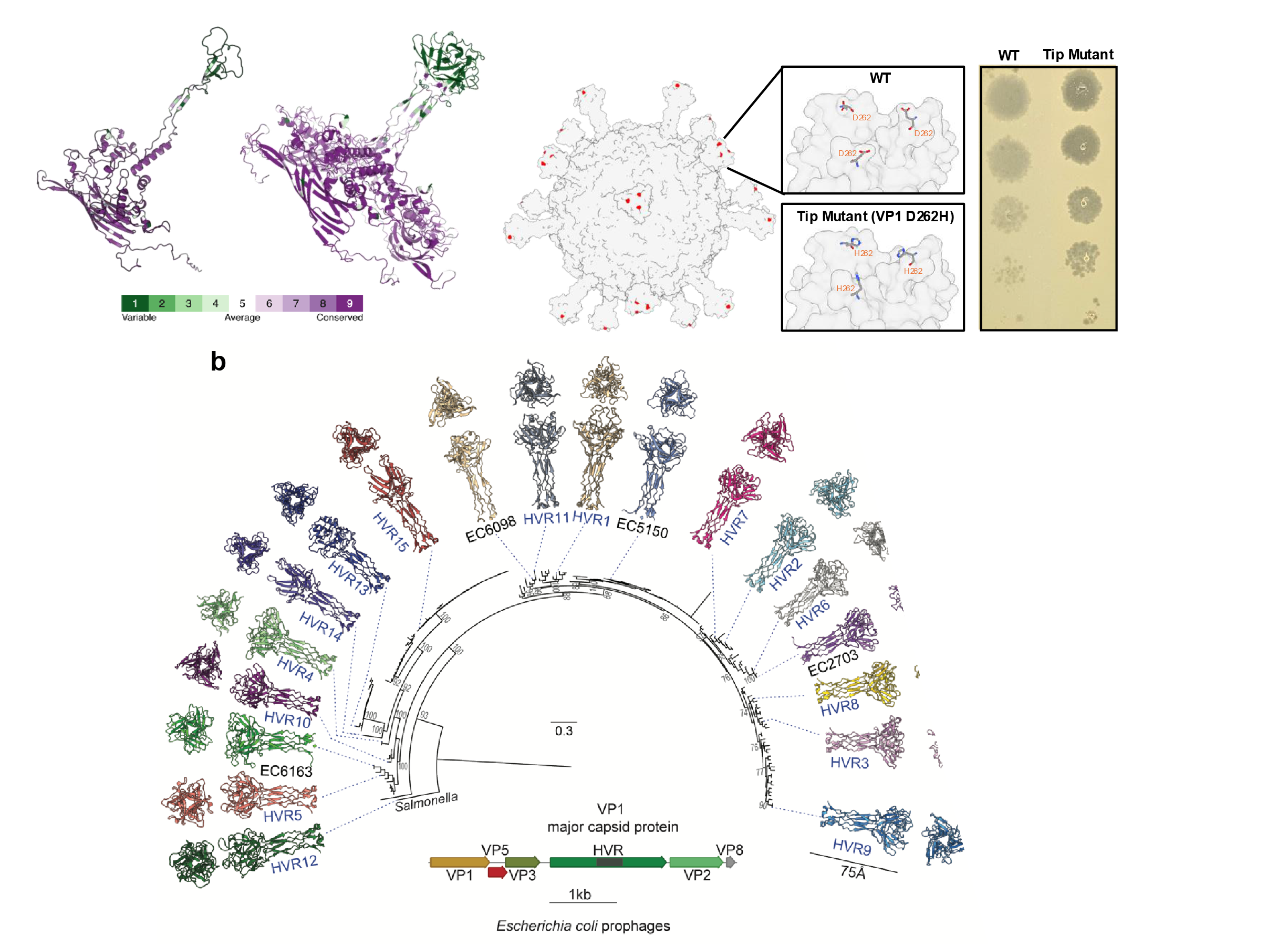
Host Range Determination
Enterogokushoviruses (and most microviruses overall) sport characteristic mushroom-like protrusions on their virions. These protrusions are formed by hypervariable regions in the capsid proteins. The regions evolve exceptionally fast, are exchanged between viruses via recombination, and can also be exchanged and manipulated in the lab.
We suspect that these regions determine a phage’s ability to infect different bacterial strains and are key to targeting phages towards specific targets. Using a combination of targeted alteration, random mutagenesis and evolution of both viral and bacterial partners, we want to determine the factors involved in this process.
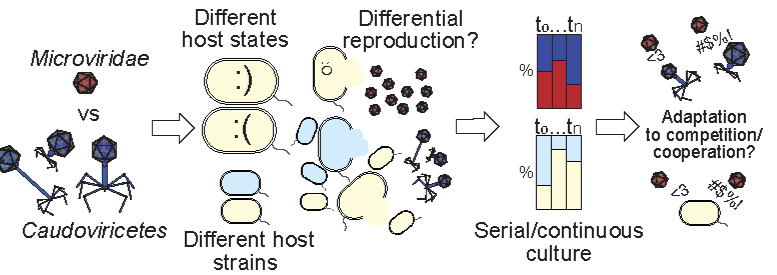
Phage-Phage Competition
Microviruses share their hosts with a plethora of unrelated, much larger phages and it remains unknown how these small viruses compete with much larger double-stranded DNA phages.
Additionally, many microvirus lineages have independently evolved the ability to integrate into the genome of bacterial hosts.
This drastic change in survival strategy has each time resulted in the concurrent evolution of defense mechanisms to protect the host from infection by other microviruses. These in turn evolve to overcome these mechanisms, creating an evolutionary arms race between infecting and defending phage.
Our immediate goals are to investigate the molecular mechanism(s) of phage competition, compare them between different microviruses, investigate the evolutionary dynamics that lead to the evolution of defenses.
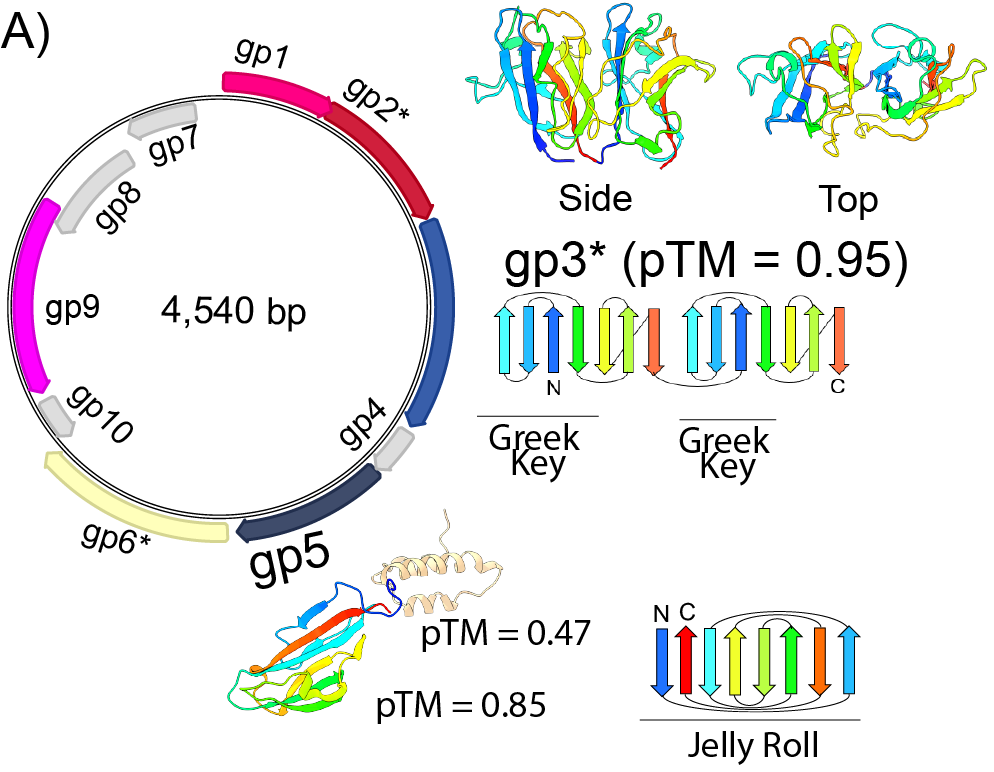
Identification, Isolation and Characterization of Novel (Micro)viruses
Metagenomic analyses indicate a world teeming with viruses, but almost none have been isolated and subject to study.
We are developing new methods based on the biology of microviruses to allow us to obtain physical isolates of these phages for comparative study.
We are also interested in “bycatch” of other non-tailed viruses that we discover along the way!